- Workflow development: Creating customized bioinformatics pipelines for diverse research goals, such as cancer genomics or microbiome studies.
- Data interpretation: Translating raw sequencing data into meaningful biological insights.
- Variant analysis: Identifying genetic variants, including single nucleotide polymorphisms (SNPs) and structural variations.
- Transcriptomics (RNA-Seq): Analyzing gene expression patterns to understand biological regulation
Overview of VBC’s NGS data analysis service
Our common service package for NGS big data analysis
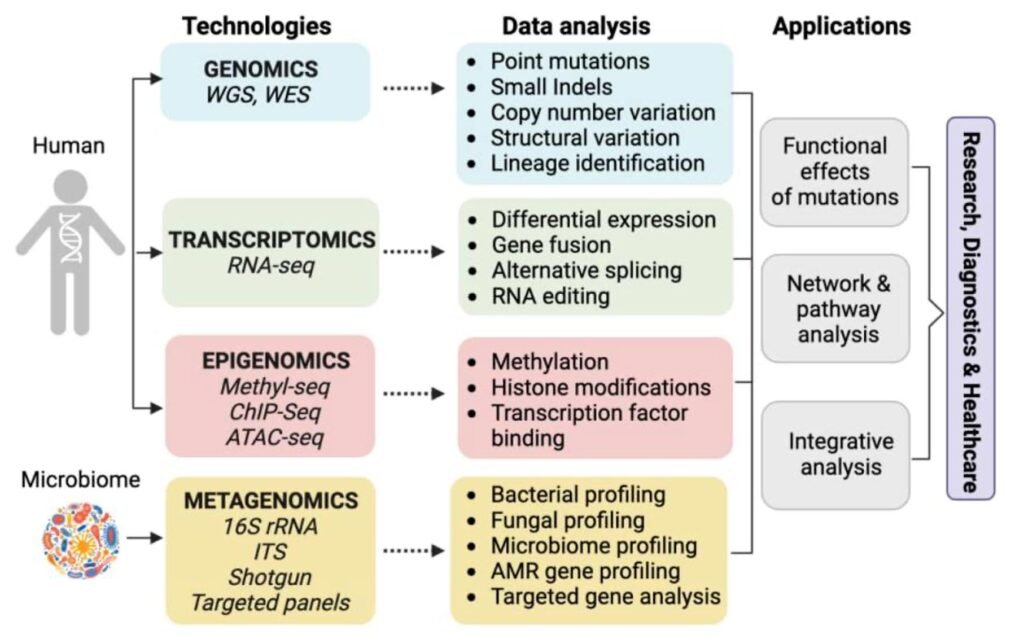
1. Genomic DNA Sequencing Analysis
Genomic DNA Sequencing Analysis using Next-Generation Sequencing (NGS) is a high-throughput technology that sequences millions of DNA fragments simultaneously, revolutionizing genomics by rapidly determining nucleotide orders to uncover genetic variations, understand disease mechanisms (like cancer), drive personalized medicine, and analyze complex biological systems, involving sample prep, library construction, parallel sequencing, and advanced bioinformatics for mapping reads to a reference genome. It enables broad, deep analysis of whole genomes, exomes, or targeted regions, detecting single-nucleotide variants (SNVs), structural changes, and gene expression patterns far beyond traditional methods.
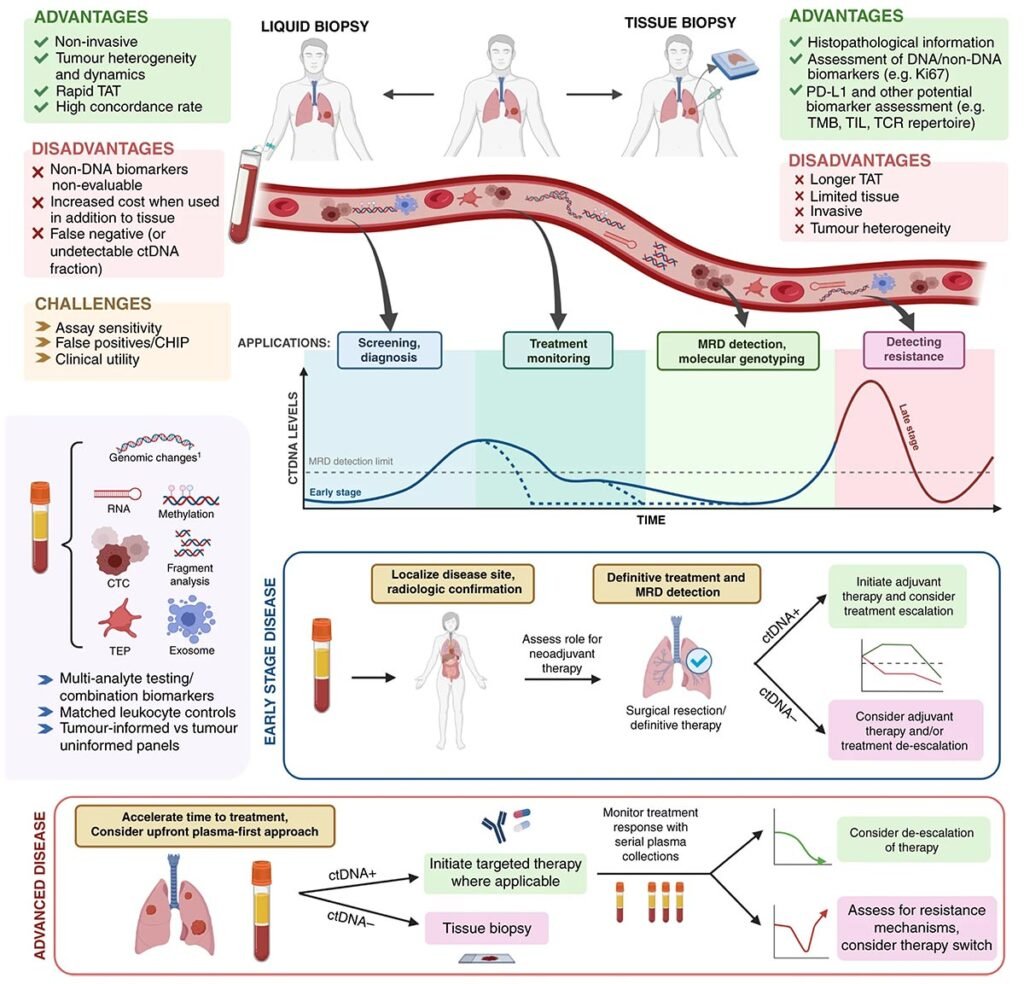
2. Circulating tumor DNA, ctDNA for Somatic Variant Detection
Circulating tumor DNA (ctDNA) analysis using Next-Generation Sequencing (NGS) is a powerful, non-invasive method (liquid biopsy) used in oncology to detect and monitor cancer-specific genetic alterations. It involves sequencing fragmented tumor DNA found in bodily fluids (primarily blood plasma) to provide real-time information about a tumor’s genomic landscape.
ctDNA is a small portion of cell-free DNA (cfDNA) released into the bloodstream by dying or viable tumor cells. NGS technology can sequence millions of DNA fragments simultaneously, allowing for the detection of rare tumor-specific mutations at very low frequencies (as low as 0.01% Variant Allele Frequency or VAF).
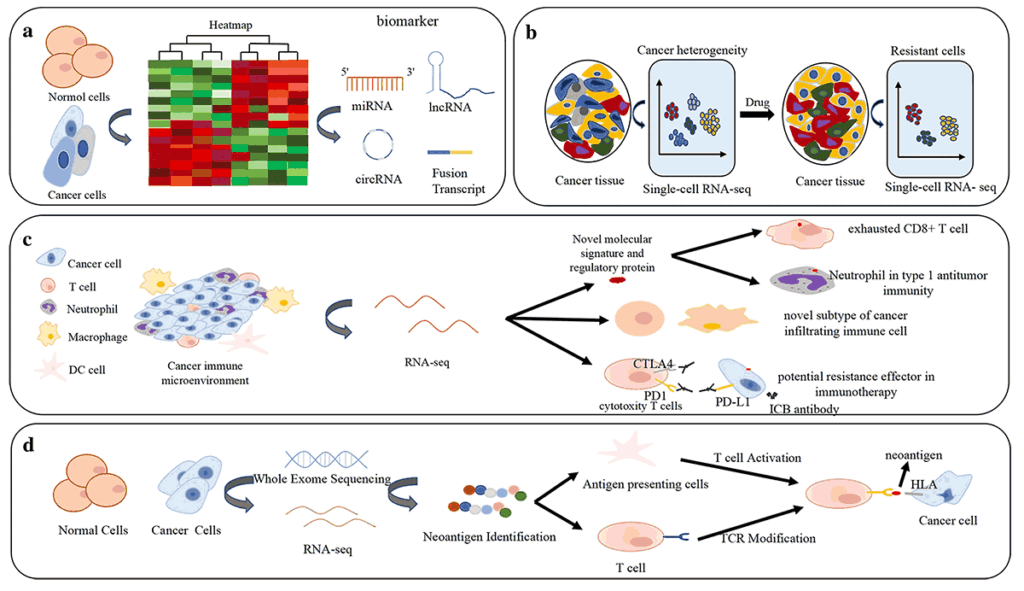
3. Gene expression profiling by RNA-Seq NGS
NGS (Next-Generation Sequencing) gene expression profiling in cancer uses RNA-Seq to create a detailed snapshot of all active genes (transcriptome) in a tumor, revealing how genes are turned on or off, identifying cancer subtypes, predicting treatment response (like to immunotherapy), finding novel biomarkers, monitoring minimal residual disease (MRD), and guiding personalized therapies, offering more depth than older methods by detecting gene fusions, alternative isoforms, and expression levels across the entire transcriptome.
RNA sequencing: new technologies and applications in cancer research
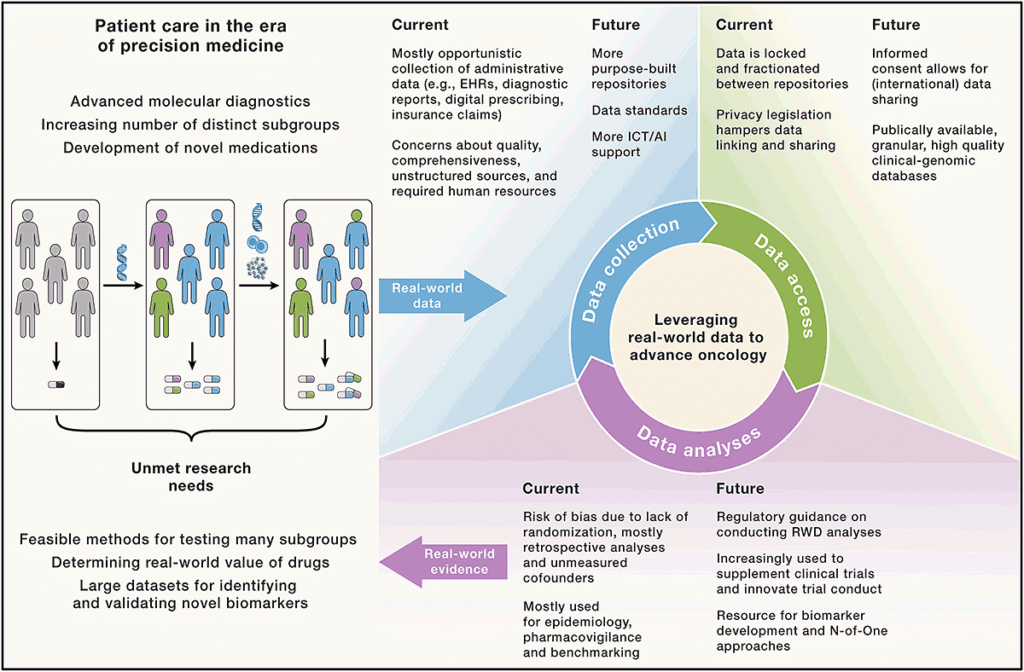
4. Real-World Data (RWD) Analysis
Real-world data (RWD) analysis of Next-Generation Sequencing (NGS) involves applying complex bioinformatics to massive datasets from routine clinical use, revealing how NGS impacts patient care, especially in oncology, by guiding targeted therapies, diagnosing rare diseases, and improving outcomes like survival, though its value depends on clinical context, with successes seen in complex cancers but limitations in rapidly progressing cases. Key analysis steps involve primary (alignment/filtering), secondary (variant calling), and tertiary (interpretation/clinical correlation) stages, utilizing cloud platforms and tools to find genomic alterations (SNVs, fusions) and link them to treatment adjustments (NBTA) for better patient management.
Combining real-world data (RWD) has enormous potential to aid in prediction of tumor trajectories.
References (as example of few previous published works developed by Peter and teams for NGS data analysis):
- Kijak et al., AIDS Res Hum Retroviruses. 2013. Nautilus: a bioinformatics package for the analysis of HIV type 1 targeted deep sequencing data (https://pubmed.ncbi.nlm.nih.gov/23809062/)
- Kijak et al., Biomol Detect Quantif 2019. Next-generation sequencing of HIV-1 single genome amplicons (https://pubmed.ncbi.nlm.nih.gov/30923677/)
- Tovanbutra et al., Cells. 2019. Deep Sequencing Reveals Central Nervous System Compartmentalization in Multiple Transmitted/Founder Virus Acute HIV-1 Infection (https://pubmed.ncbi.nlm.nih.gov/31443253/)
- Lewitus et al., mBio. 2024. HIV-1 Gag, Pol, and Env diversified with limited adaptation since the 1980s (https://pubmed.ncbi.nlm.nih.gov/38329340/
